Expression Datasets
01 Annona squamosa Pulp Ripening
RNA-Seq of Annona squamosa (cv. Gefner and African Pride) pulp at different stages of growth and ripening.
Experimental Conditions (3 biological replicates):
- African Pride pulp 30 DAP.
- African Pride pulp 100 DAP.
- African Pride pulp 130 DAP.
- African Pride pulp 6 DAH.
- African Pride pulp 8 DAH.
- Gefner pulp 30 DAP.
- Gefner pulp 100 DAP.
- Gefner pulp 130 DAP.
- Gefner pulp 6 DAH.
- Gefner pulp 8 DAH.
The samples were obtained from the "Guangxi Academy of Agricultural Sciences" in Guangxi, China. Thirty fruit samples were collected for each cultivar at five stages of growth and ripening between April and July 2019. Fruits from each cultivar and stage were separated into three biological replicates, and pulp samples were collected, frozen in liquid nitrogen, and used for RNA-seq. The sampled time points correspond to 30, 100, and 130 DAP, as well as 6 and 8 DAH.
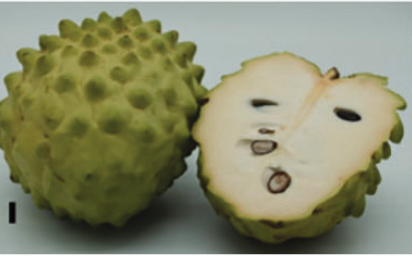









Images of fruits from A. squamosa cv. African Pride (top) and cv. Gefner (bottom) at 30, 100 and 130 DAP; and 6 and 8 DAH.
DAP = Days After Pollination
DAH = Days After Harvest
This dataset was published by Fang et al. 2020, and raw data can be found in the BioProject PRJNA639613.
02 Annona squamosa Flower Development
RNA-Seq of Annona squamosa (cv. Bendi) flowers at different stages of development.
Experimental Conditions:
- Inflorescence meristeme.
- Flower bud.
- Mature flower with partially opened petals.
- Mature flower with opened and faded petals.
The samples were obtained from the "Lingnan Normal University" in Guangdong, China. The inflorescence meristem (IM), the flower buds (FB), and two stages of flowers (FL1 and FL2) were collected from A. squamosa trees. The flower buds were collected based on their size (3–6 mm). The two flower stages were the mature flowers with partially opened petals (FL1) and mature flowers with opened and faded petals (FL2). These tissue samples were frozen immediately in liquid nitrogen, and stored at −80 ºC until use. Equal volumes of RNA from each of the four stages of flower development were pooled, and each sample was prepared by mixing three replicate samples.
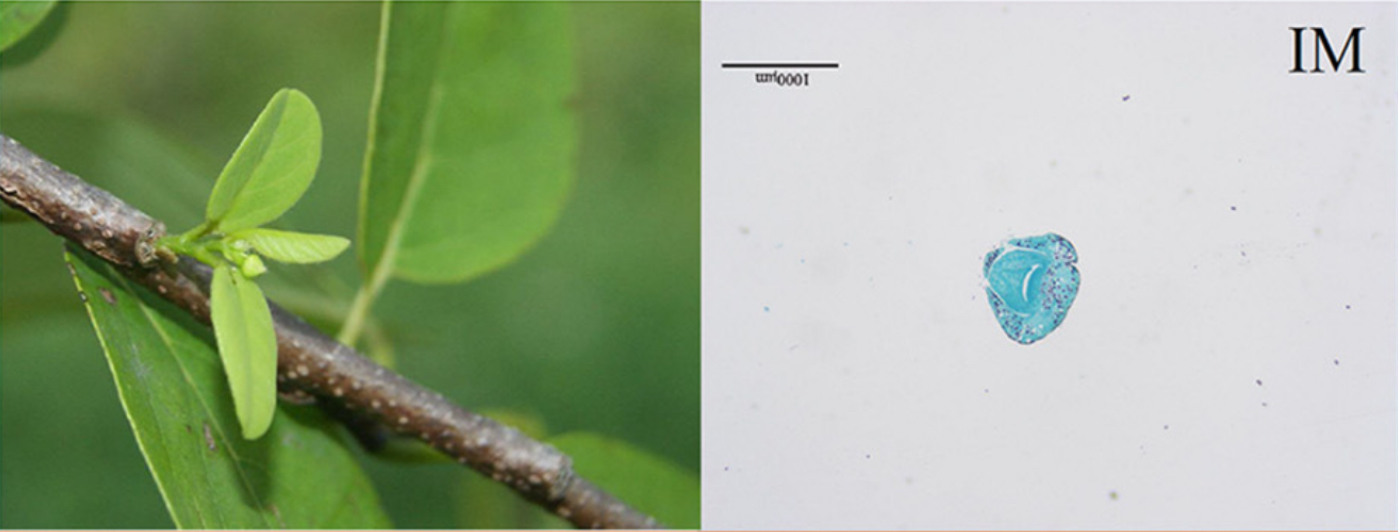

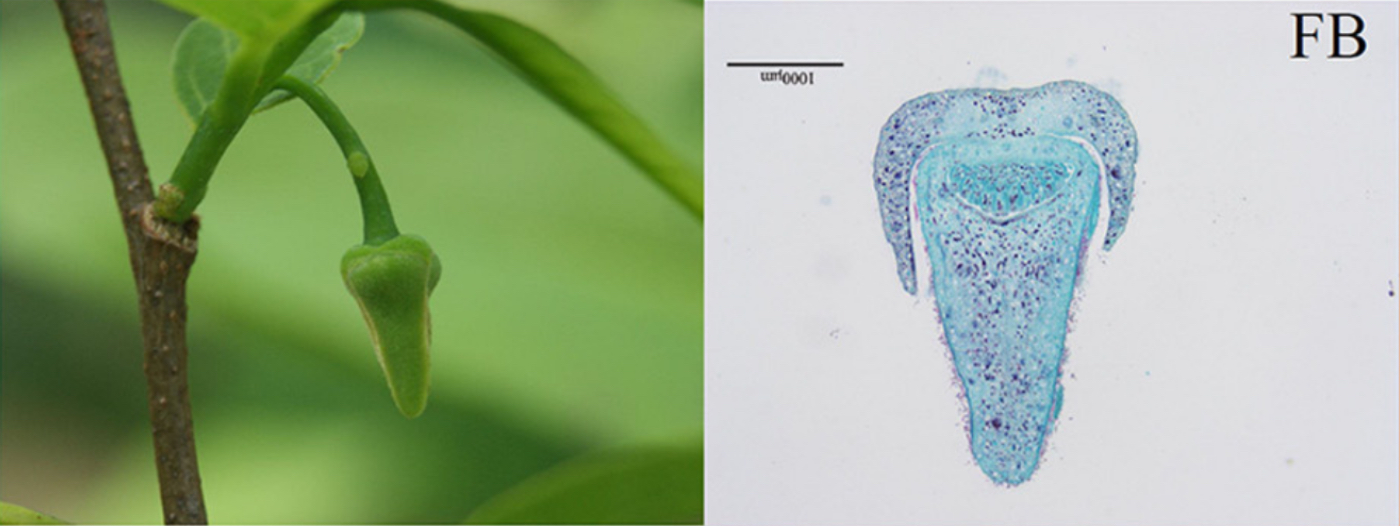
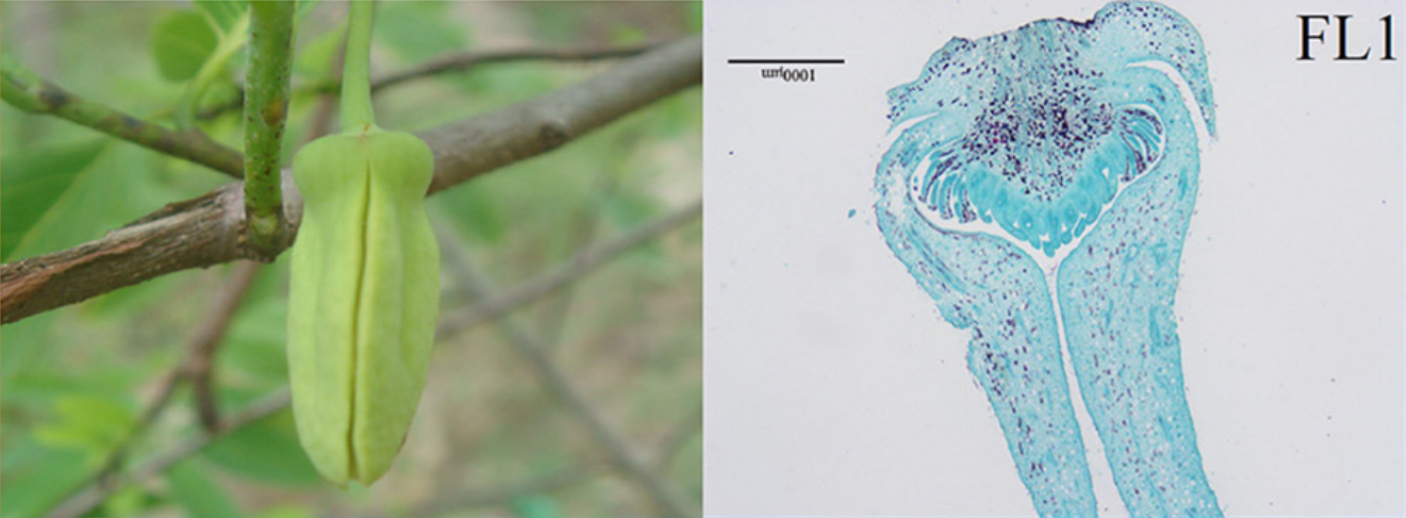
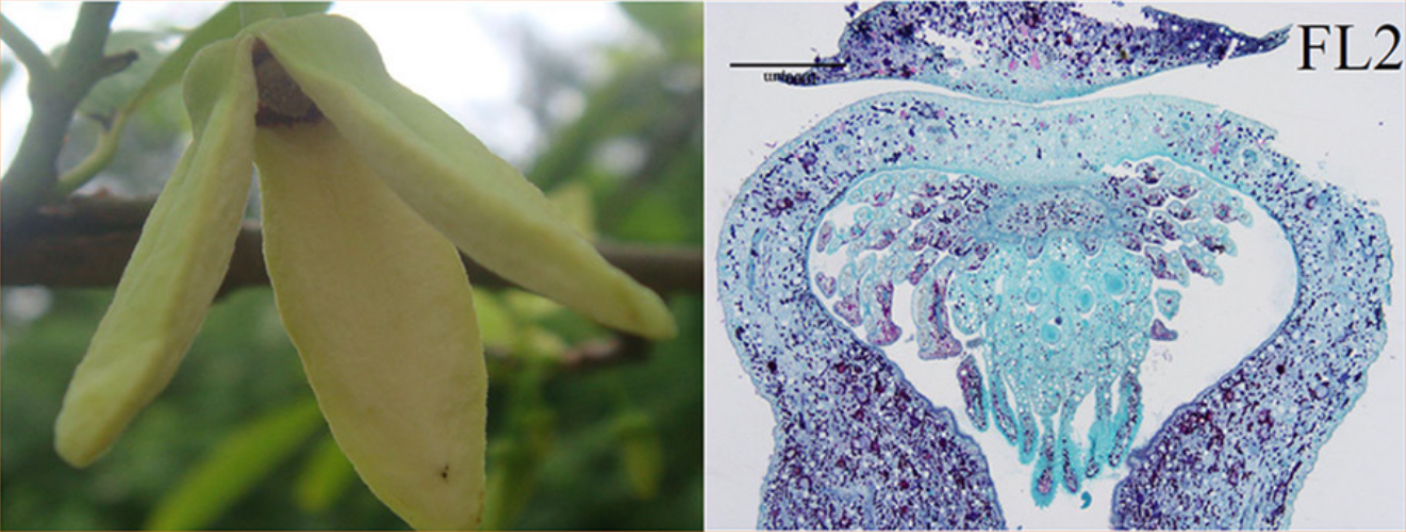
Images of flowers from A. squamosa (cv. Bendi).
IM = Inflorescence Meristeme
FB = Flower Bud
FL1 = Mature flower with partially opened petals
FL2 = Mature flower with opened and faded petals
This dataset was published by Liu et al. 2016, and raw data can be found in the BioProject PRJNA320668.
03 Annona squamosa Normal and Malformed Flowers
RNA-Seq of Annona squamosa (cv. Bendi) normal flowers and malformed flowers.
Experimental Conditions:
- Normal flowers.
- Malformed flowers.
The samples were obtained from the "Lingnan Normal University" in Guangdong, China. The flowers before open were collected from a normal flower and a malformed flower from A. squamosa trees. All of the flower samples were frozen immediately in liquid nitrogen, and stored at −80 ºC until use. Each sample of RNA was prepared by mixing three replicate samples.
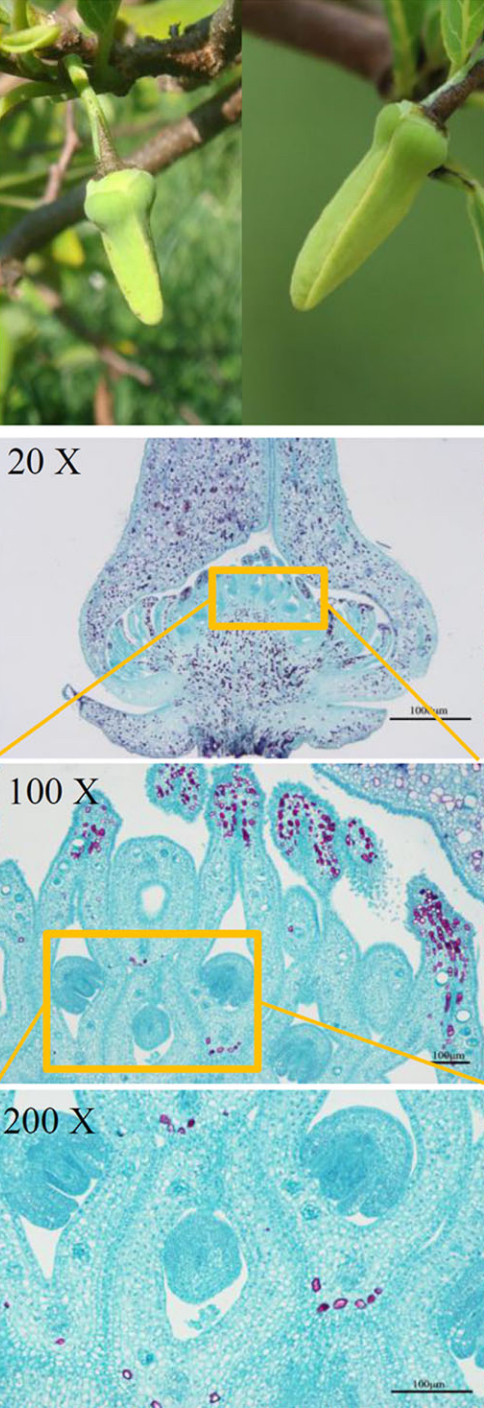
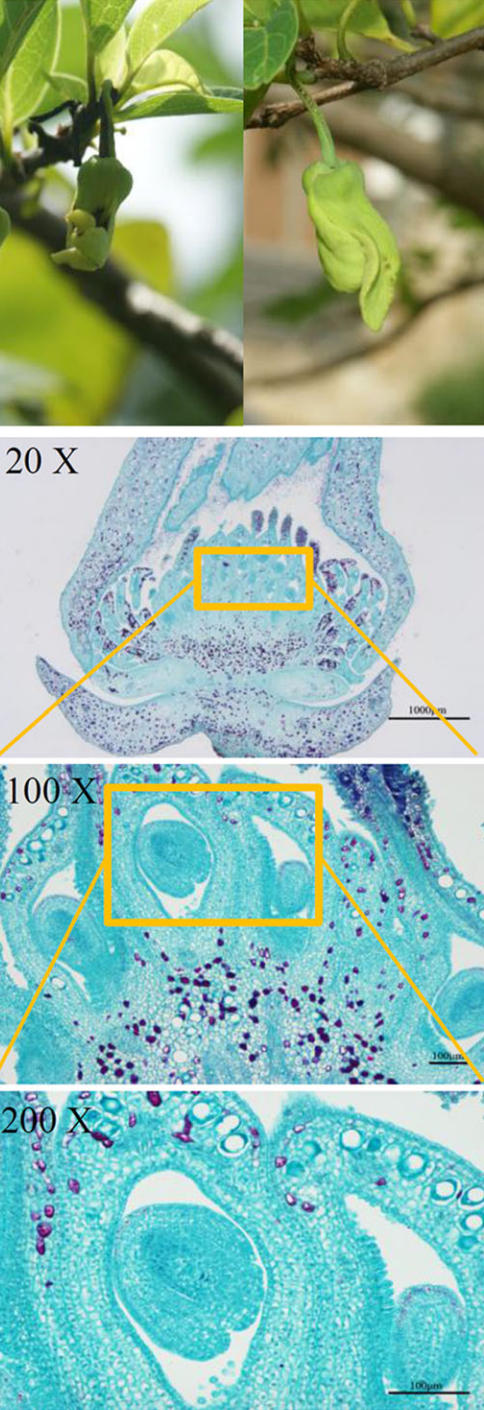
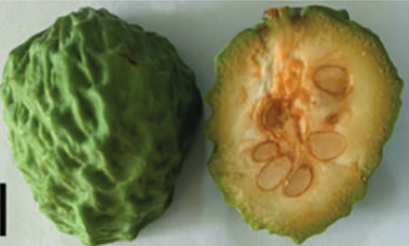
Images of normal flowers (left) and malformed flowers (right) from A. squamosa (cv. Bendi).
This dataset was published by Liu et al. 2017, and raw data can be found in the BioProject PRJNA358456.
04 Atemoya Pericarp Fruits with Ethylene Treatment
RNA-Seq of atemoya (Annona squamosa × A. cherimola) (cv. African Pride) pericarp fruits during post-harvest storage with and without ethylene treatment.
Experimental Conditions:
- Normal treated fruits.
- Ethylene treated fruits.
The samples were obtained from the "Chinese Academy of Tropical Agricultural Sciences" in Guangdong, China. Twenty fruits were kept at room temperature with no treatment, and the following four stages were selected for transcriptome sequencing: recently picked fruits (NT-0), cracking at the pedicel (NT-PC-1), moderate cracking at the pedicel (NT-PC-2), and severe cracking at the pedicel (NT-PC-3). Another twenty fruits were immersed in 2 g/kg ethylene for 2 min, transferred into a closed plastic box, and the lid was left open after 24 hours. The following three stages were selected for transcriptome sequencing: fruits 24 hours after ethylene treatment (Eth-24), fruits with pericarp cracking after ethylene treatment (Eth-PC), and no fruit cracking after ethylene treatment (Eth-NC).
Three similar pericarp samples under the same treatment at each stage were collected, immediately frozen in liquid nitrogen, and then stored at −80 °C until use. Each sample of RNA was prepared by mixing three replicate samples.
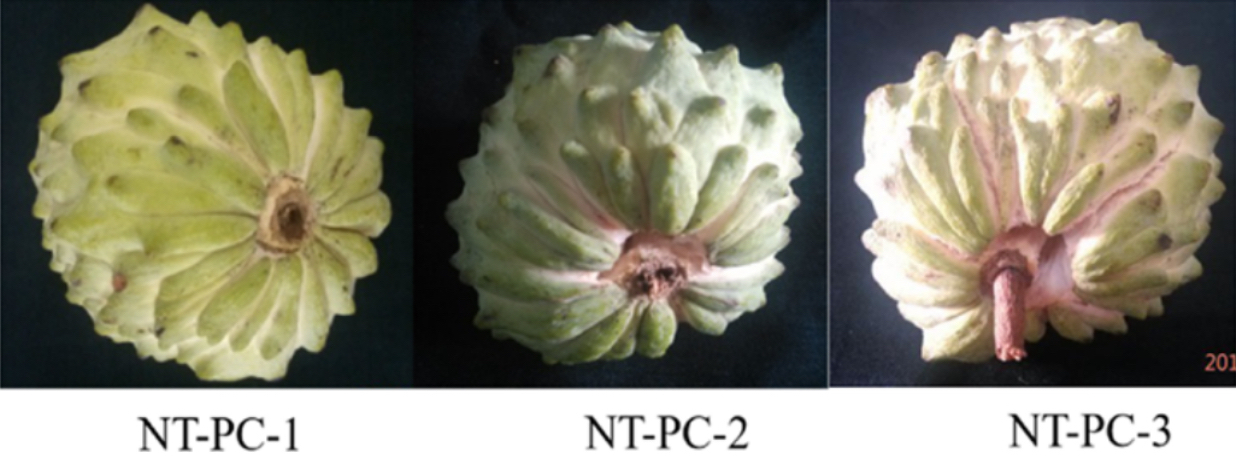


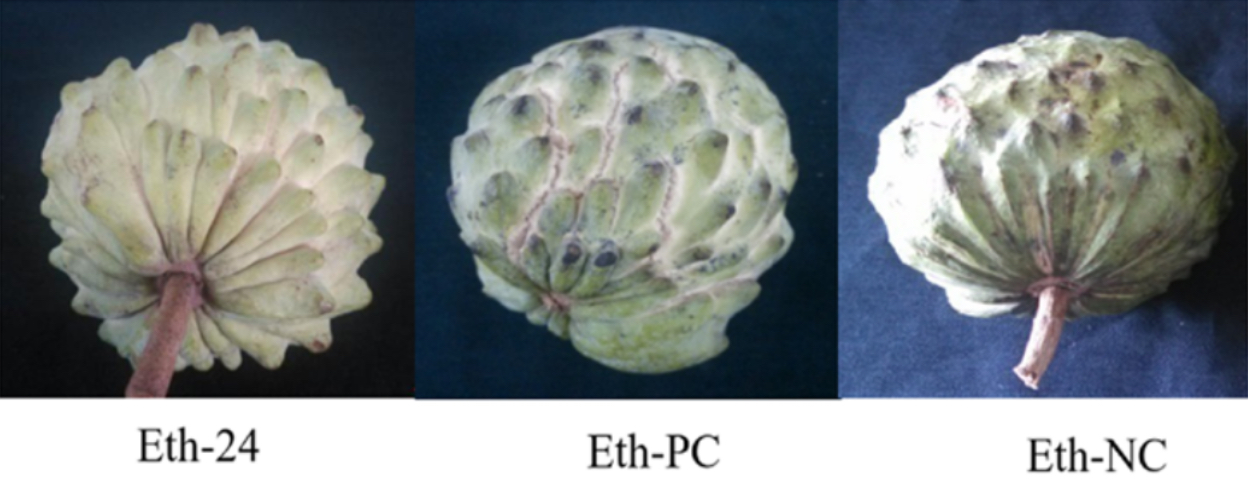
Images of fruits from atemoya (cv. Africa Pride) during ripening and cracking in normal conditions and treated with ethylene.
NT = Normal Treatment
PC = Pericarp Cracking
Eth = Ethylene Treatment
NC = No Cracking
This dataset was published by Chen et al. 2019, and raw data can be found in the BioProject PRJNA396469.
05 Atemoya Split and Non-Split Fruits
RNA-Seq of atemoya (Annona squamosa × A. cherimola) pulp of split and non-split fruits.
Experimental Conditions:
- Split fruits.
- Non-split fruits.
The samples were obtained from the "Lingnan Normal University" in Guangdong, China. All fruits were harvested at 6 months after pollination. Non-split fruits (NSF) and split fruits (SF) were selected at 10 days after harvest. All of the fruit samples were frozen immediately in liquid nitrogen, and stored at −80 ºC until use. Each sample of RNA was prepared by mixing three replicate samples.
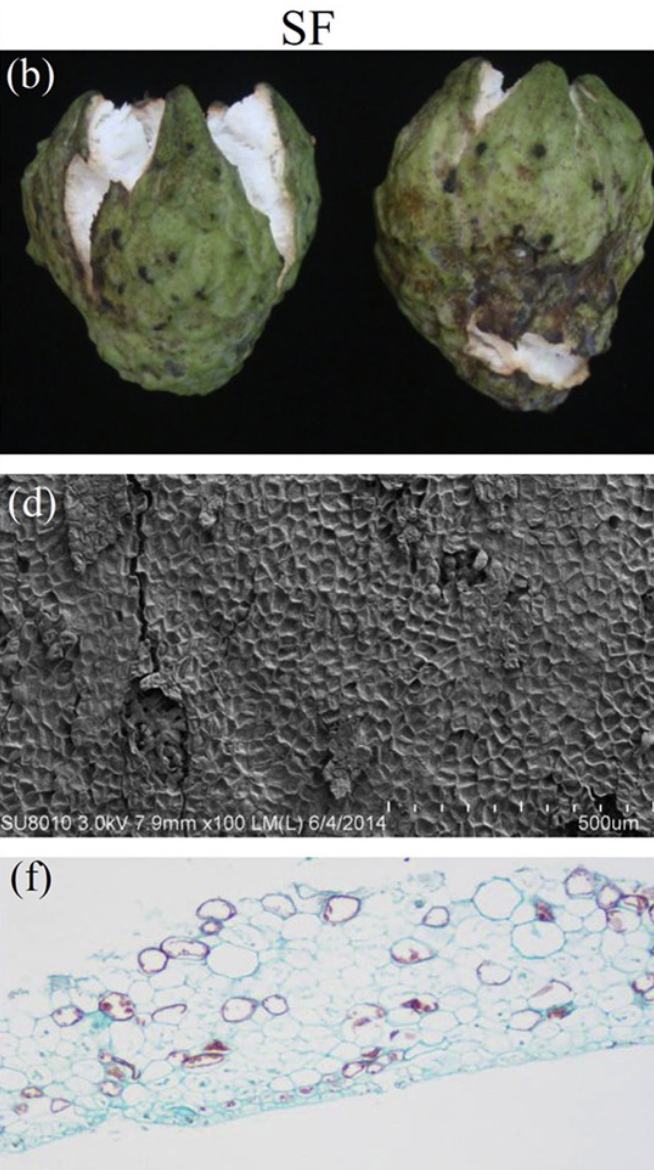
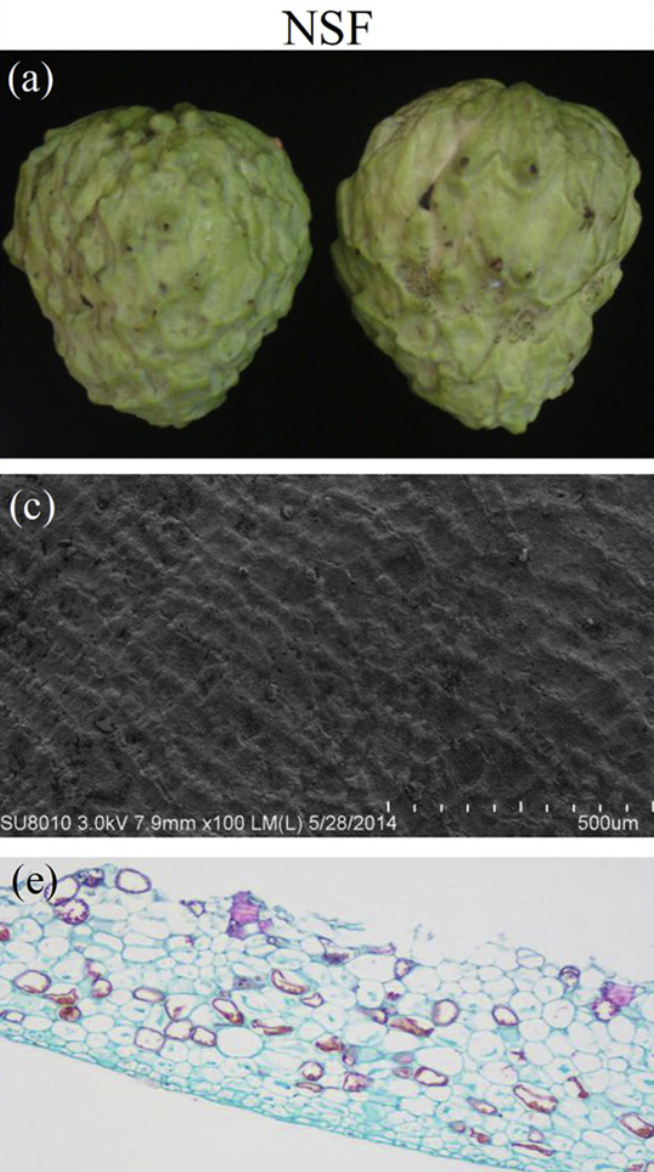


Images of NSF (a) and SF (b) from atemoya. Observation with Scanning Electron Microscopy (SEM) the NSF (c) and SF (d) in atemoya, and observation with Optical Microscope the NSF (e) and SF (f).
NSF = Non-Split Fruits
SF = Split Fruits
This dataset was published by Li et al. 2019, and raw data can be found in the BioProject PRJNA397365.
06 Annona Outer Integument
RNA-Seq of pistils with and without early seed development in Atemoya (Annona squamosa × A. cherimola) cv. Gefner, and the A. squamosa "Thai seedless" mutant (Annona ino).
Experimental Conditions:
- AG = Atemoya cv. Gefner
- T = "Thai seedless" mutant
- PS = Pistils containing early developing seeds (Pollinated pistils with seed).
- P = Pistils without early developing seeds (Pollinated pistils, seed removed).
Total RNA was extracted from pollinated pistils containing early seeds (PS) and pistils without early seeds (P) of Annona cherimola cv. Gefner (AG) and the "Thai seedless" mutant (T), combining the CTAB protocol by Doyle and Doyle, 1987 and the Qiagen RNeasy Plant Mini Kit (Qiagen).
PS and P samples were collected from three independent fruits at 4 and 8 days after pollination (DAP).
Microscopic observations were performed at 8 DAP to illustrate early embryo and nucellus development.
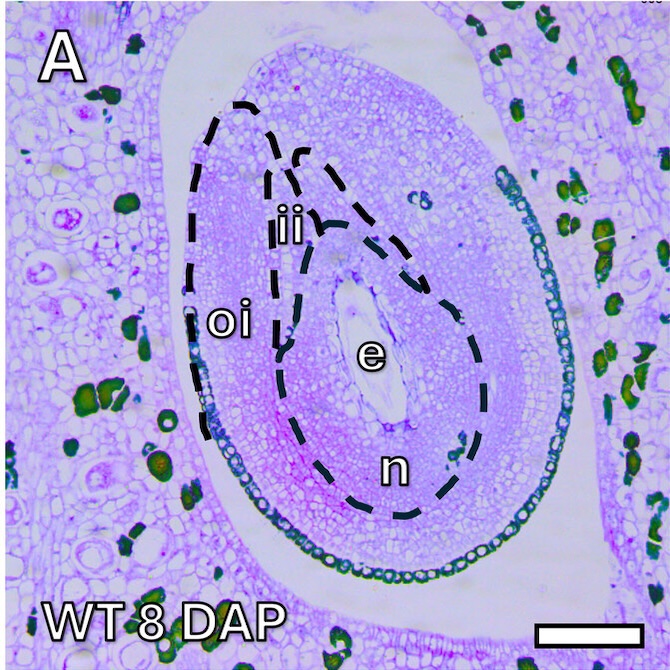
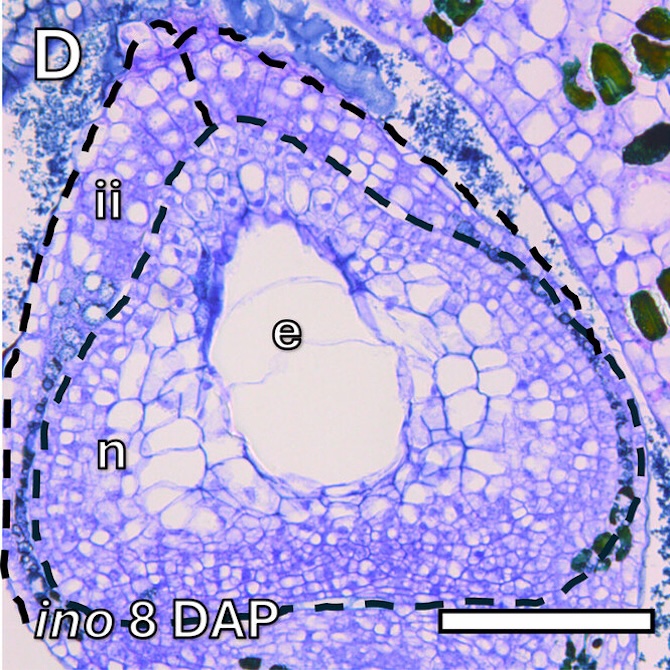
Microscope images of the immature seed sections (8 DAP) for both AG (A) and T (D) genotypes.
e = endosperm
ii = inner integument
n = nucellus
oi = outer integument
Scale bars: 100 μm
This dataset was published by Garcia-Lezama et al., 2025 in Physiologia Plantarum. Raw sequencing data are available in the BioProject PRJEB97455.
Asimina triloba reproductive incompatibility
RNA-Seq of pawpaw (Asimina triloba) early developing seeds collected from three cross-compatible (CC) and three auto-incompatible (AI) pollinated flowers at 4, 8 and 15 days after pollination (DAP), resulting in a total of 18 biological replicates. The pistils of each flower were dissected to obtain the developing seeds, and extract RNA from these fresh early seeds.
Experimental Conditions:
- Developing seeds of cross-compatible pollinated flowers at 4 DAP (CC4)
- Developing seeds of auto-incompatible pollinated flowers at 4 DAP (AI4)
- Developing seeds of cross-compatible pollinated flowers at 8 DAP (CC8)
- Developing seeds of auto-incompatible pollinated flowers at 8 DAP (AI8)
- Developing seeds of cross-compatible pollinated flowers at 15 DAP (CC15)
- Developing seeds of auto-incompatible pollinated flowers at 15 DAP (AI15)
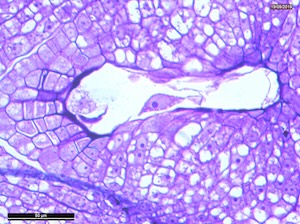
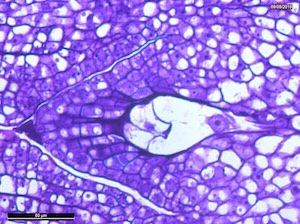 4 DAP
4 DAP
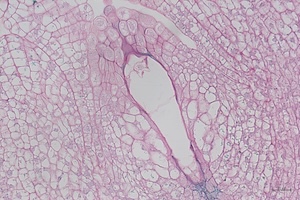
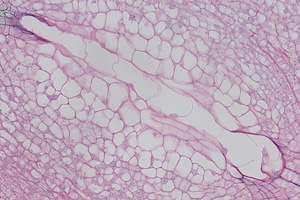 8 DAP
8 DAP
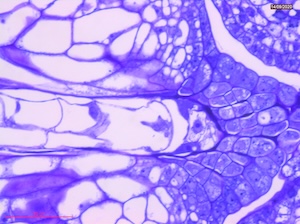
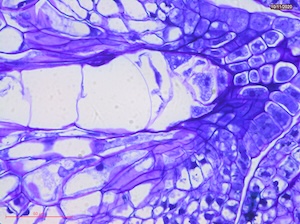 15 DAP
15 DAP
Microscope images of developing seeds of cross-compatible (on the left) and auto-incompatible crosses (on the right).
CC = compatible crosses
AI = Autoincompatible crosses
DAP = Days After Pollination
This dataset was published by Ferrer-Blanco et al. 2025, and raw data can be found in the BioProject PRJEB97068.
T.cacao Phytophthora infection
RNA-Seq of pods of seven cacao genotypes (4 susceptible: ICS1, WFT, Gu133, Spa9, and 3 resistant: CCN51, Sca6 and Pound7) to study their response to the post-penetration stage of Phytophthora palmivora infection.
| Variety | Genetic Background / Characteristics |
|---|---|
| Pound7 | Upper Amazon Forastero lineage; known for specific disease resistance traits. |
| Sca6 | Scavina 6; a foundational clone for Crinkly Leaf and Black Pod resistance. |
| CCN51 | Colección Castro Naranjal; a high-yielding, widely cultivated commercial hybrid. |
| Spa9 | Selection from the 'Servicio de Plantas Agrícolas' (Ecuador). |
| Gu133 | Guiana selection; representing wild genetic diversity from French Guiana. |
| WFT | Specific selection used for pathology and transcriptomic modeling. |
| ICS1 | Imperial College Selection 1; a high-quality Trinitario clone. |
Experimental Conditions (3 replicates):
- Control Group: Healthy pod tissues, non-infected.
- Infection Group: Pod tissues infected with Phytophthora palmivora, the primary causal agent of Black Pod disease.
Methodology & Bioinformatics Pipeline:
- Library Generation: RNA-Seq libraries were generated specifically from cacao pod RNA.
- Platform: Sequencing was conducted on an Illumina platform.
- Mapping: Reads were aligned to the reference Matina v2 genome.
- Quantification: Gene expression levels were normalized using TPM (Transcripts Per Million) values to allow for accurate comparative analysis across genotypes and treatments.
This dataset was published by Baruah et al. 2024 and raw data can be found in the BioProject PRJNA487154.
01 C. papaya fruit ripening induced by ethylene
RNA-seq of Carica papaya (cv. "Golden") samples in five groups of analysis, depending on the treatment and the ripening time.
Experimental conditions (3 biological replicates):
- C0h
- C12h
- C24h
- T12h
- T24h
The samples were obtained from the “Department of Food Science and Experimental Nutrition, School of Pharmaceutical Sciences, University of São Paulo”. Papayas (Carica papaya L. cv. “Golden”) at the pre-climacteric stage were acquired from a commercial producer in the municipality of Linhares/ES, Brazil, between one and two days after harvest, still with up to one-quarter of yellow peel (about 150 days after anthesis) and were harvested in three consecutive harvesting times (August, September, October 2017). Right after the fruit arrived in the laboratory and following sanitization with chlorine (100 ppm), some were immediately characterized and frozen to compose the 0-h control group. Remaining fruits were separated into two groups: treated and control. Control groups were left to ripen, and fruit samples were taken after 12 h and 24 h. Treatment with ethylene at 0 h was done by exposing randomly selected fruit to a concentration of 100 ppm (100 µLL−1) of ethylene, kept in constant flux for 17 min for gas saturation and 12 h more in a closed system. After 12 h, fruits were removed from the ethylene chamber and exhaustively air-vented for 1 h. Ethylene-treated fruits were frozen (12 h group), and the other set of fruit was left to ripen as the control group for 12 more hours, totaling 24 h after fruits reached the laboratory (group 24 h treated). In all five groups (0 h, 12 h control, 24 h control, 12 h ethylene, and 24 h ethylene), papayas were characterized for ripening parameters and peeled; their seeds were removed, and the pulp was cut into cubes, frozen in liquid nitrogen, and stored at −80 ◦C. The experiment used three sets of harvested fruit collected at three different times of the year (biological triplicate), and each group was composed of at least five fruits (which were pooled before storage at −80 ◦C).
- C = Control.
- T = Treated. Randomly selected fruit exposed to a concentration of 100 ppm (100 µLL−1) of ethylene.
This dataset was published by Soares et al., 2021, and raw data can be found in the PioProject PRJNA528193.
02 C. papaya fruit ripening of two varieties with different flavor profile
RNA-seq of Carica papaya varieties (RB1 and 1B) samples at ripe and unripe stages.
Experimental conditions (3 biological replicates):
- 1B ripe stage
- 1B unripe stage
- RB1 ripe stage
- RB1 unripe stage
The samples were obtained from the “Griffith University”. Two Australian commercial cultivars, RB1 and 1B, were harvested at ripening stages 1 and 3, based on the previously developed harvest ripening index, from a commercial papaya plantation in Mareeba, in June 2021. Three fruits of each variety at each fruit maturity stage were used as biological replicates. A total of 12 fruit samples were treated with Sportak to control postharvest disease, stored at 27 ◦C for one day and then transported in a refrigerated truck to the laboratory at Griffith University, Nathan Campus, Brisbane. Fruit flesh was cut into 5 mm pieces and flash frozen in liquid N2, then stored at −80 ◦C until processed for RNA extraction.
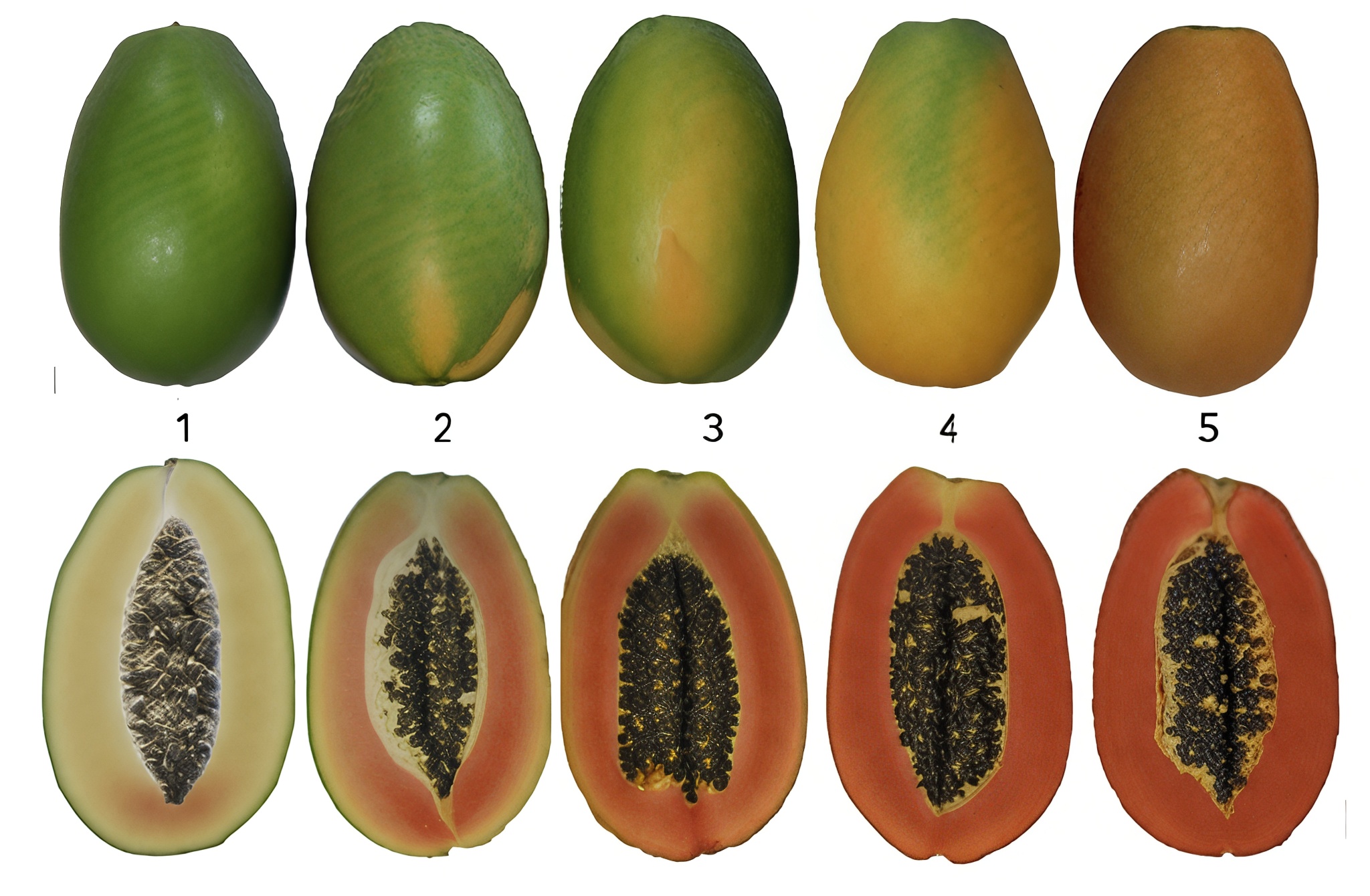

Five fruit maturity and ripening stages (1 = mature green, 2 = 25% colour, 3 = 50% colour, 4 = 75% colour, 5 = fully ripe).
- 1B = yellow-fleshed cultivar
- RB1 = red-fleshed cultivar
This dataset was published by Zhou et al., 2024, and raw data can be found in the PioProject PRJNA880626.
03 C. papaya pistil development
RNA-seq of Carica papaya carpel on different development stages.
Experimental conditions (3 biological replicates):
- Female S1
- Male S1
- Female S2
- Male S2
- Female S3
- Male S3
- Female S4
- Male S4
The samples were obtained from the “Fujian Agriculture and Forestry University”. Female and male flowers of papaya were sampled and fixed according to flower size and length and stored in FAA liquid. After putting samples through an alcohol dehydration series, flower samples were embedded in paraffin (LEICA EG 1150 H) in a Leica EG 1150 H paraffin embedding apparatus (Leica, Germany) at a melting point of 50-52 °C. Transverse sections with a thickness of 10 μm were made on a Leica automatic rotary microtome and stained with Brilliant Green (BIO-RAD). The sectioned and embedded papaya flowers were dissected under a dissecting microscope (Olympus, Japan) to isolate the different developmental stages of the male and female papaya flowers. Due to the difficulty in sampling pistils from male and female flowers at the primordium stage, the flower buds at the primordium stage was used as the experimental material for this study.
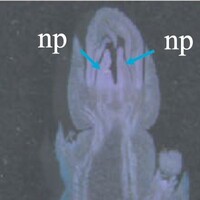
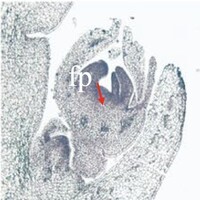
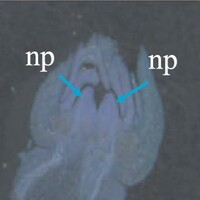

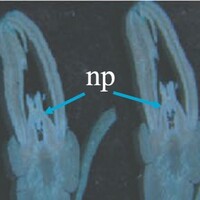
The phenotype of the papaya pistil at different developmental stages (S1 to S4) in female flowers.
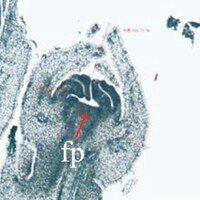
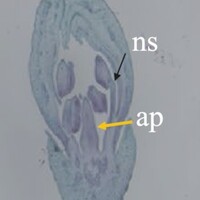
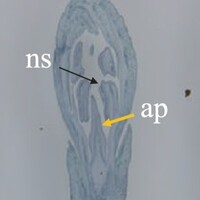
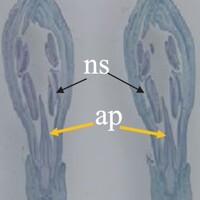
The phenotype of the papaya pistil at different developmental stages (S1 to S4) in male flowers.
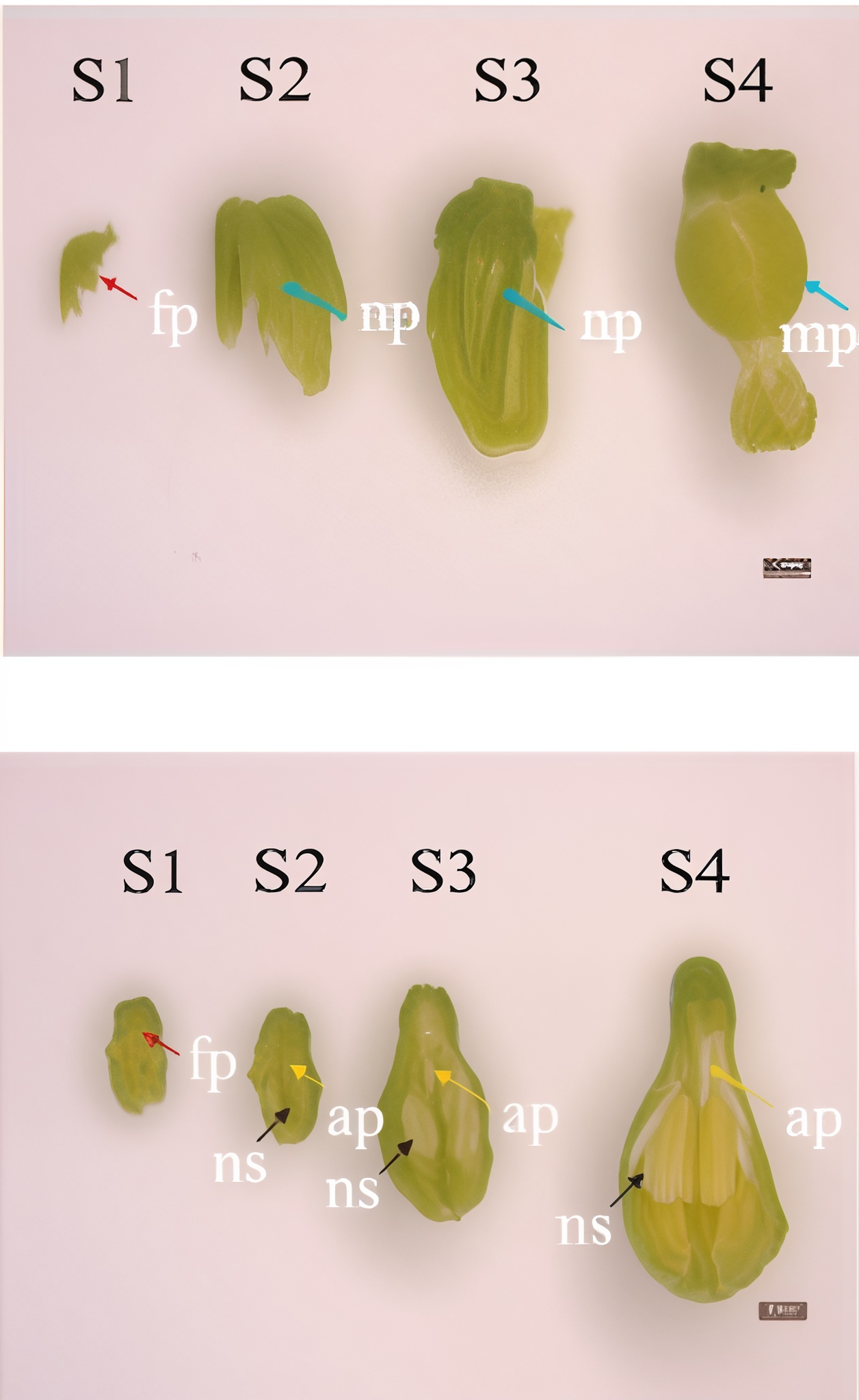

The phenotype of the papaya pistil at different developmental stages (S1 to S4) in female and male flowers.
- S1 = primordium stage
- S2 = pre-meiotic stages
- S3 = post-meiotic stage
- S4 = mitotic stage
- fp = floral primordium
- np = normal pistil
- ap = aborted pistil
- ns = normal stamen
This dataset was published by Liao et al., 2022, and raw data can be found in the PioProject PRJNA687615.
04 C. papaya flowers buds at pre- and post-meiosis
RNA-seq of Carica papaya (var. AU9 and SunUp) samples among three different sex types, during early (pre-meiosis) and later (post-meiosis) stages of flower development.
Experimental conditions (2 biological replicates):
- AU9F.1
- AU9F.2
- AU9M.1
- AU9M.2
- SUH.1
- SUH.2
The samples were obtained from the “University of Illinois at Urbana-Champaign”. Flower buds were collected from female and male ‘AU9’ papaya plants and hermaphrodite ‘SunUp’ plants grown at the Kunia Research Station of Hawaii Agriculture Research Center (HARC) in 2013. The flower buds were used to compare gene expression between sex types and obtain candidate sex determination genes by RNA-Seq. These flower buds were first classified according to their phenotype (sex) and then were divided into two groups according to their size (in millimeters). Flower buds were ground in liquid nitrogen for further RNA extraction. To further corroborate the differential expression of identified highly differentially expressed genes by qPCR, flower buds, and leaf tissue samples were collected again from three different ‘SunUp’ female plants, three different ‘SunUp’ hermaphrodite plants, three different ‘AU9’ female plants and three different ‘AU9’ male plants grown at the Kunia Research Station of HARC during 2017. All samples were collected in Hawaii by HARC, shipped in dry ice (−80C) to Urbana, Illinois and then ground in liquid nitrogen (− 196C) for further RNA extraction.
- AU9F.1 = Female flower bud size between 1 and 6 mm, “AU9” cultivar.
- AU9F.2 = Female flower bud size between 7 and 12 mm, “AU9” cultivar.
- AU9M.1 = Male flower bud size between 1 and 6 mm, “AU9” cultivar.
- AU9M.2 = Male flower bud size between 7 and 12 mm, “AU9” cultivar.
- AU9F.1 = Hermaphrodite flower bud size between 1 and 6 mm, “SunUp” cultivar.
- AU9F.2 = Hermaphrodite flower bud size between 7 and 12 mm, “SunUp” cultivar.
This dataset was published by Zerpa-Catanho et al., 2019, and raw data can be found in the PioProject PRJNA565901.
05 C. papaya gynoecium morphogenesis induced by auxin
RNA-seq of Carica papaya (var. Zhongbai and Zhonghuang) gynoecius samples from different flowers (hermaphrodite, male and female).
Experimental conditions (3 biological replicates):
- H_dom-FG
- H_aux-RG
- H_dom-RG
- F-FG
- F.FG
- M.MG
The samples were obtained from the “Fujian Agriculture and Forestry University”. Gynodioecious papaya variety “Zhongbai” and dioecious papaya variety “Zhonghuang” were planted in the research station of Fujian Agriculture and Forestry University (FAFU). In November 2015, small inflorescences including rudimentary gynoecium of hermaphroditic dominant and auxiliary flowers, functional gynoecium of hermaphroditic dominant and female flowers from “Zhongbai”, as well as, functional gynoecium of female flowers and rudimentary gynoecium of male flowers from “Zhonghuang” were collected near the trunk apex and preserved in RNA later solution for 2 days. The ambient temperature fluctuated from 20 ˚C at night to 31 ˚C at day for these two days. To investigate the differentially expressed genes between rudimentary and functional gynoecium, five comparisons were designed. Four comparisons were from “Zhongbai” includes the functional gynoecium of hermaphroditic dominant flowers and rudimentary gynoecium of auxiliary flowers, functional gynoecium of hermaphroditic dominant flowers and rudimentary gynoecium of hermaphroditic dominant flowers, functional gynoecium of female flower and rudimentary gynoecium of hermaphroditic dominant flowers, and functional gynoecium of female flower and rudimentary gynoecium of hermaphroditic auxiliary flowers, while, one was from dioecy variety “Zhonghuang” (functional gynoecium of female flower and rudimentary gynoecium of male flower). We selected early-stage flowers with length <2 mm and separated functional and rudimentary gynoecium by anatomic dissection under microscope for RNA extraction. Three biological replications for each type of sample were collected with each replication of 5–30 flowers.
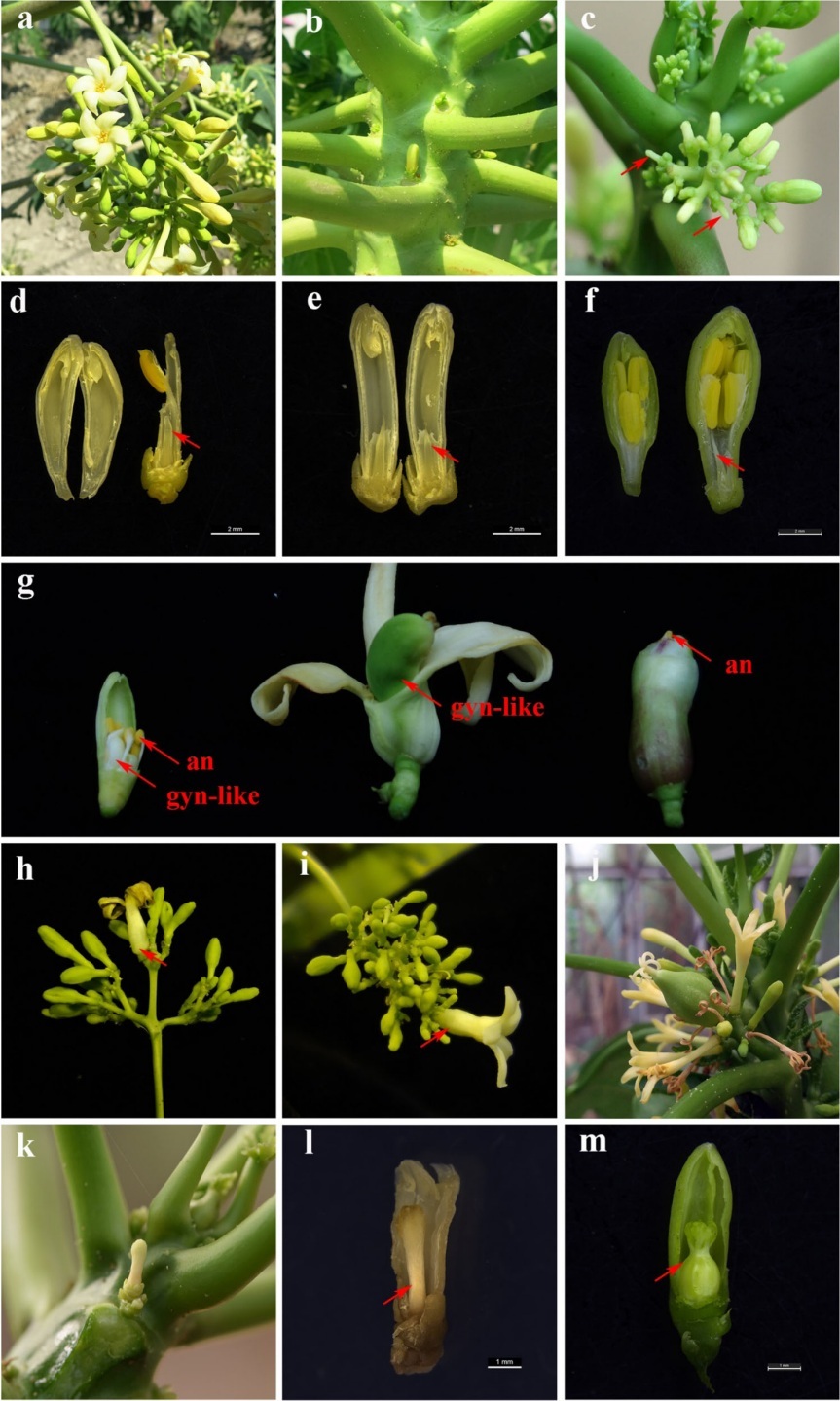

The phenotypic changes of the NPA-treated flowers in dioecious papaya “Zhonghuang”. (a) Male inflorescence on the male plants. (b) Female flowers on the female plants. (c) Male inflorescences treated by NPA. Arrow indicated a long-rod phenotype flowers due to a decrease in androecium number. (d) A long-rod phenotype male flower with only one androecium. Arrow indicated the rudimentary gynoecium. Bar= 2 mm. (e) A long-rod phenotype male flower with no trace of filaments and anthers. Arrow indicated the rudimentary gynoecium. Bar=2mm (f) Normal male flower. Arrow indicated the rudimentary gynoecium. Bar =2 mm. (g) Phenotypically similar perfect hermaphrodite flowers with fertile gynoecium in male plant. Gynoecium-like structures (gyn-like) bore flat and smooth stigmas(left), bore feathery stigmas (mid), and entirely embedded in merged outer whorls of floral system(right). All newly induced hermaphrodite flowers referred to have visible androecium(an). (h and i) The transformed primary male flowers surrounded with adjacent clusters of male flowers. Arrow indicated the transformed primary male flowers. (j) The inflorescences of Vasconcellea monica. (k and l) The NPA-treated female flower bore a stalk-like gynoecium with stigma. Arrow indicated the stalk-like gynoecium, Bar= 1mm (m) Normal female flower. Arrow indicated the functional gynoecium, Bar =1mm
- H_dom-FG = Hermaphrodite dominant flowers, functional gynoecius, fertile, var. Zhongbai
- H_aux-RG = Hermaphrodite auxiliary flowers, rudimentary gynoecius, sterile, var. Zhongbai
- H_dom-RG = Hermaphrodite dominant flowers, rudimentary gynoecius, sterile, var. Zhongbai
- F-FG = Female flowers, functional gynoecius, fertile, var. Zhongbai
- F.FG = Female flowers, functional gynoecius, fertile, var. Zhonghuang
- M.RG = Male flowers, rudimentary gynoecius, sterile, var. Zhonghuang
This dataset was published by Zhou el al. 2019, and raw data can be found in the PioProject PRJNA549650.
06 C. papaya callus and shoot regeneration
RNA-seq of Carica papaya (var. Zhongbai) samples at five different timepoints: hypocotyl, callus 1, callus 2, callus 3, and shoot.
Experimental conditions (3 biological replicates):
- Hypocotyl
- Callus 1
- Callus 2
- Callus 3
- Shoot
The samples were obtained from the “Fujian Agriculture and Forestry University” (FAFU). The papaya cultivar ‘Zhongbai’ from FAFU was used in this study. All samples were grown in greenhouse of FAFU at a temperature of 28 ± 2 ◦C and 16 h light/8 h dark photoperiod. The light intensity is set to 100 μmol/m2 /s. Papaya seeds were surface sterilized using 75 % ethanol for 1 min, 3.5 % sodium hypochlorite solution for 20 min and 0.1 % mercuric chloride for 10 min and washed three times with distilled water, then soaked in 3.5 % sodium hypochlorite solution for 4 h (150 rpm at 28 ◦C). Seeds were then soaked in 1.0 M KNO3 for 24 h (100 rpm at 28 ◦C) and then transferred to distilled water until germination (100 rpm at 28 ◦C).





Morphological observation of hypocotyl, callus (1, 2 and 3) and shoot during in vitro organogenesis of papaya.
- Hypocotyl = 7 days on MS
- Callus 1 = 21 days on CIM K5
- Callus 1 = 21 days on CIM M13
- Callus 1 = 21 days on CI
- Shoot = 28 days on SIM
This dataset was published by Zhao et al., 2021, and raw data can be found in the PioProject PRJNA713527.
07 C. papaya drought stress
RNA-seq of Carica papaya (cv. Maradol Roja) different tissues (leaves, sap, and roots) under well-watered condition and drought stress.
Experimental conditions (2 biological replicates):
- Leaf CN
- Sap CN
- Root CN
- Leaf 10 DASI
- Sap 10 DASI
- Root 10 DASI
- Leaf 20 DASI
- Sap 20 DASI
- Root 20 DASI
The samples were obtained from the “Yucatan Center for Scientific Research”. After germination, the papaya plants were grown under greenhouse conditions and equivalently watered. Drought stress was imposed on three-month-old plants by withholding watering; healthy well-watered plants of same age were used as controls. Visual assessment of phenotype, physiological measurements, and sample collection for RNA isolation, were performed on plants under control condition and at 10 and 20 days after stress imposition.



Three-month-old ‘Maradol’ papaya plants under control condition.



Three-month-old ‘Maradol’ papaya plants under 10 days after stress imposition.



Three-month-old ‘Maradol’ papaya plants under 20 days after stress imposition.
- CN = under control condition (well-watered)
- DASI = days after stress imposition
This dataset was published by Gamboa-Tuz, S.D., Pereira-Santana, A., Zamora-Briseño, J.A. et al. 2018, and raw data can be found in the PioProject PRJNA470602.
08 C. papaya response to PLDMV and Chitosan-N
RNA-seq of Carica papaya (cv. No.2 in Tai Nong) leaves in three different conditions.
Experimental conditions (3 biological replicates):
- B group
- CK group
- CG group
The samples were obtained from the “Hainan University”. The experiment was divided into three groups: the treatment group, the negative control group, and the positive control group. Leaves of the treatment and positive control groups were inoculated with PLDMV. The treatment group was administered 1 mL/L CTS-N as well, and the negative control measured normal natural growth. Leaves were ground evenly and placed in a −80 °C freezer.
- B = treatment group
- CK = negative control group
- CG = positive control group
This dataset was published by An et al., 2020, and raw data can be found in the PioProject PRJNA591254.


